Whole Genome Sequencing - 101
by Péter Szilágyi
The “science” of longevity is living its renaissance. Countless doctors, nutritionists and even influencers are preaching the best way to achieve your longest lifespan and best health-span. They have everything figured out, ranging from the perfect exercise routine, through the perfect diet and supplementation, down to the exact protocol for even less-understood substances.
The problem is, it’s not easy to find two, who agree on all those details. We all have that one friend who swears on the carnivore diet; but also the other one, who never ate meat in their life. We have that friend who does crossfit five times per week, and the other whose peak workout is the trip from the bed to the computer - yet is fit as ever. Thing is, each of us is unique and what might work for some, could possibly fail for another.
Sure, there are generally accepted axioms that certain things are clearly good (exercise) while certain other things are clearly bad (smoking). But the lines quickly start to blur once you go beyond the basics.
The age-old solution is “try it and go with whatever works for you”. Whilst nice in theory, there are too many hidden variables that might only surface when it gets unpleasant - or worse. If we want to talk about longevity as a “science”, throwing things at the wall until something sticks is not ideal. It’s not like we have infinite time to do it the dumb way.
To “do science” (i.e. understand a phenomenon), we need to define variables we can track across experiments, so we might better understand how certain action influences the outcome. Some of these variables might be feeble and change on a whim; others might be stable, even constant across time. Most, however, will exhibit an intricate interplay.
From an engineering perspective, creating a stable foundation first is a prerequisite for building on top. Thus, before starting to juggle dynamic variables, we should define the static ones. From a biological standpoint, the strongest stable-point we have is our genes (mostly1).
Crash course on genetics
Choose your own adventure! Since not everyone’s going to be on the same page with regard to the genetics, here’s a choice for you.
-
For absolute beginners, unfortunately this post might be out of reach. The world of genetics is amazing, but immense. We strongly recommend everyone to gain at least a basic understanding of the subject. There are countless courses for free. ¯\_(ツ)_/¯
-
For novice readers, to whom it might have been a while since they last dealt with the topic of genetics, here’s a crash course to brush up on the notions. It’s definitely not for the faint of heart!
-
For advanced readers, we’re betting you don’t know the link between a pint of semen and the discovery of DNA! The lecture below will broaden your horizon while the novices brush up.
With the basics elegantly handwaved out of the way, it’s time to dive a bit deeper into the discussion. If we ever hope to understand ourselves better, we need to understand our starting point in life first, which is the strand of genetic information inherited from our parents. The only way to do that, is to actually pull the genetic code out of our bodies into a computer. The process is called DNA sequencing.
Genome vs. exome sequencing
When talking about DNA sequencing, two predominant variants come up: “whole genome sequencing” and “whole exome sequencing”. For the record, “targeted sequencing” also exists, where only a very small part of the DNA strand is analyzed. The goal there is clinical cost-efficiency, but it is of limited use outside targeted diagnostics.
Back to genome vs. exome sequencing. The difference is which parts of a DNA strand they decipher. The genome is the whole thing, sequencing that gets you everything. That said, the regions of a genome that actually encode the proteins - the exons (EXpressed regiONS) - are <2% of the DNA. The rest are introns (INTRagenic regiONS), and are discarded during transcription.
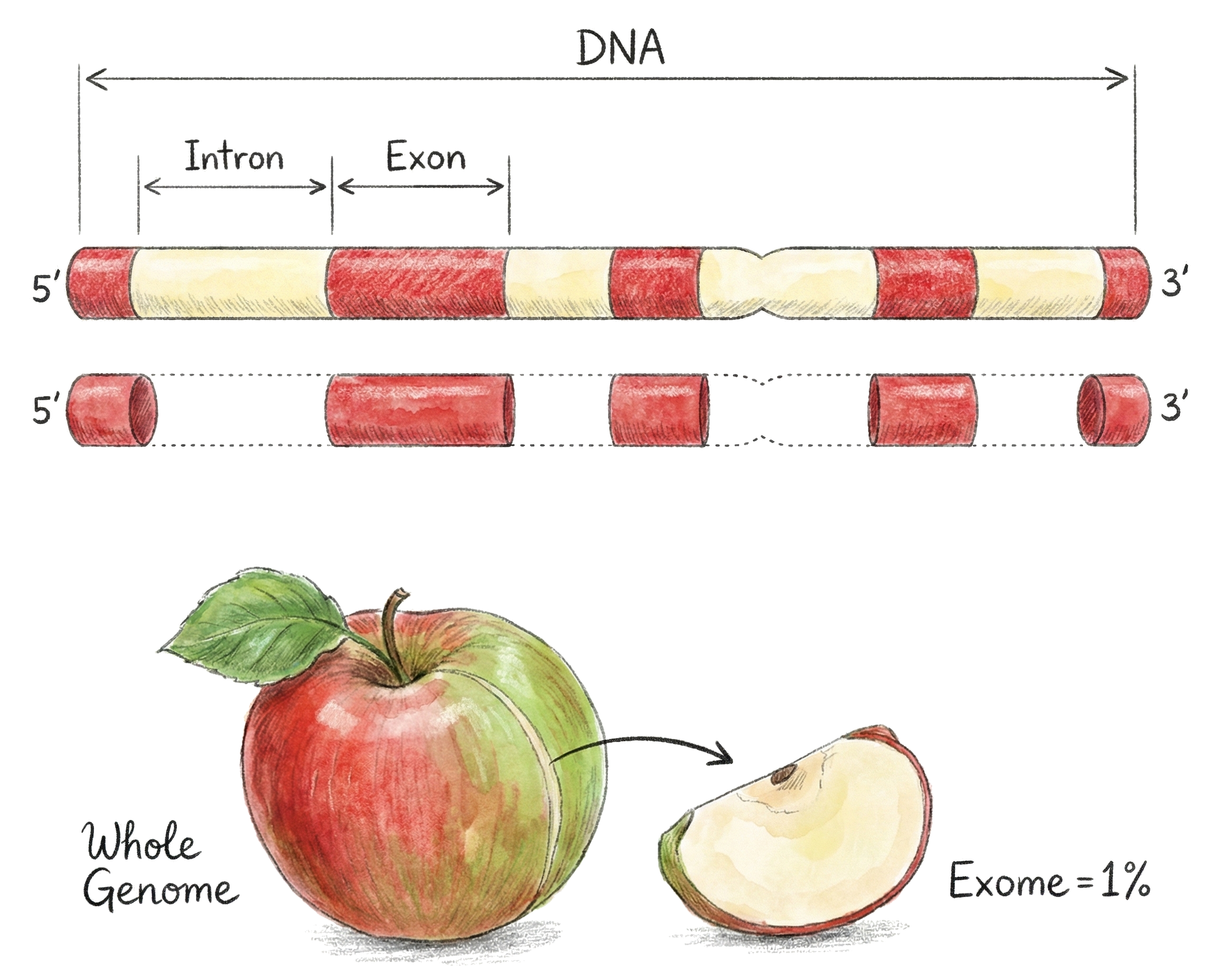
Given than exons can be extracted and sequenced cheaper, faster and result in a very compact representation of our genes, it would seem a no-brainer to sequence that. Unfortunately, while true that the introns are discarded from the final code that gets “assembled”, they themselves turn out to have a fairly large impact on how the assembly happens.
Think of it like a compiler. The DNA is the source code to build. The exons are the actual instructions to turn into the final program. But the source code is littered full with introns, which can act as pragma statements for the compiler itself to ever so slightly modify its behavior. Looking at the set of instructions will allow detecting glaring errors, but to understand the thing as a whole, the full source code is needed.
Introns have some other cute uses too like drastically raising the possible recombination space during inheritance and storing idle mutations that pile until a combination activates. Both are essential parts of evolution as the former tends to increase the chance of random mutations, while the latter locks them in via some evolutionary advantage2. We’ll ignore these for now.
Long story short, “whole exome sequencing” is useful for quickly detecting variations in a genetic code (compared to the population norm); analysing a specific individual, however, is better done via “whole genome sequencing”, where all the data is laid bare to go through - as it functions in reality. Whether we understand it or not is a whole different question.
Next generation sequencing
In the early days, the Sanger sequencing was the way to parse the DNA into a machine consumable state. It produced highly accurate results, but at an excruciatingly slow pace (originally 60KB/h, later 600KB/h, but the human genome is about 3GB). It was too expensive to use on a global scale.
Nowadays, labs are using “next generation sequencing”, which is an umbrella term for all the newer tech that came after Sanger. The brave naming is due to all the new methods3 focusing on parallel sequencing. Instead of reading a single DNA strand, they cut it up into tiny pieces and read them parallel to one another.
Being able to read many strands at once means, that these mechanisms can afford to sacrifice accuracy. If reading a strand with some non-negligible error rate is extremely fast and cheap, then those errors can be detected by just reading the DNA more than once. These sequencing methods actually start at 30x coverage and go up to over 100x to filter out inaccuracies.
The catch with massively parallel sequencing is that the length of the DNA strands they read is very small, on the order a hundred base pairs. Certain machines can do long reads, but their accuracy falls. This poses a notable challenge when assembling a genome, as it requires piecing together a 3+GB long, seemingly random sequence of data, out of tiny chunks, many of which have faults in them.
E.g. On the image below you can see a bunch of intact DNA strands (blood, saliva, etc). These get blown apart into many tiny pieces via a chemical process, and then sequenced via next generation machines. To reconstruct the original DNA strand, all the tiny pieces are overlapped. The problem lies in the sheer number of these tiny pieces.
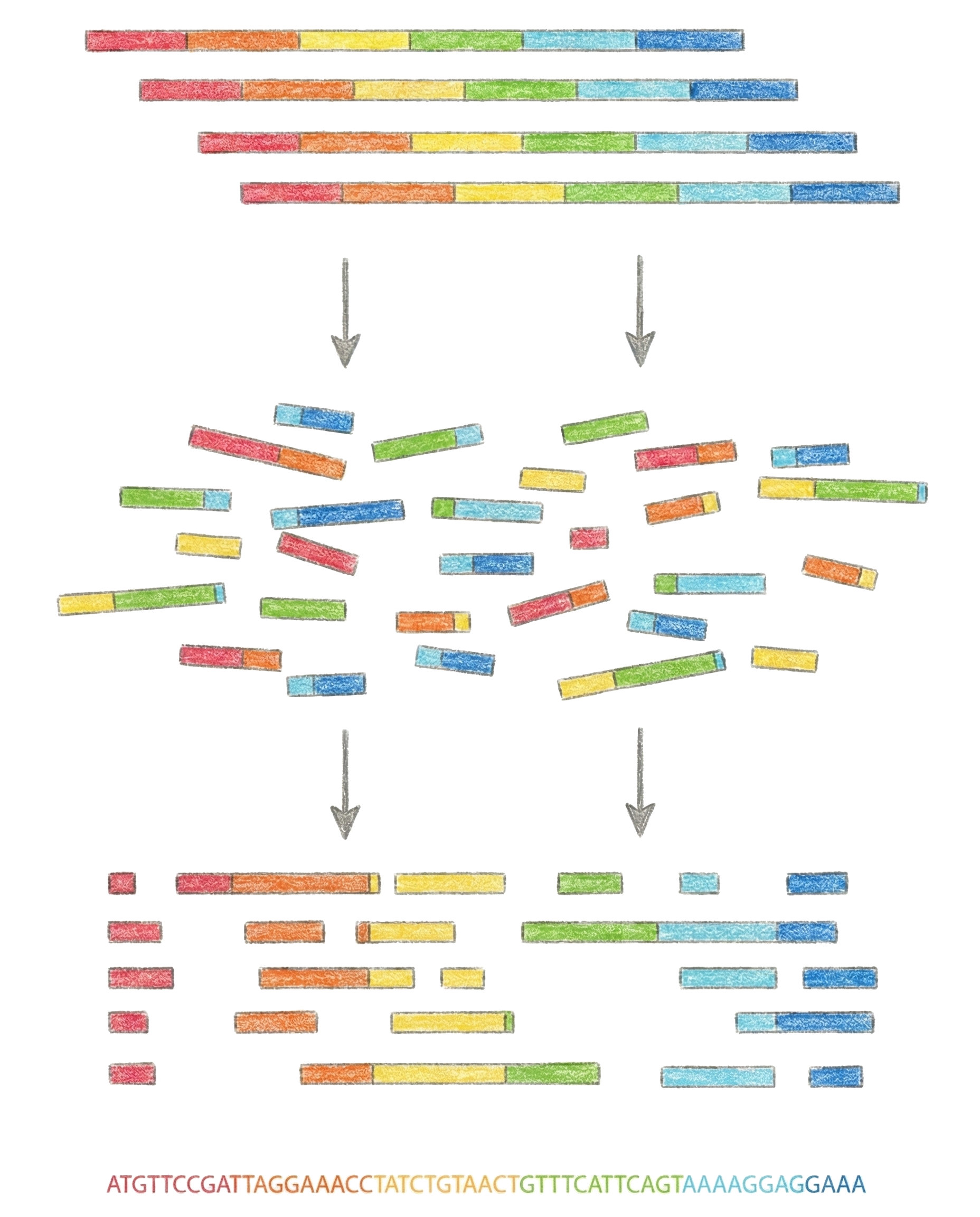
The solution is a second tradeoff. Whilst everyone’s DNA is unique, most of it is actually extremely similar between people, only tiny differences here and there. If we have a few genomes already sequenced, mapping a new person becomes a lot easier as the short chunks can be compared to a few reference datasets to see where they fit best. This is called “reference mapping”. If sequencing is done without a reference, we would be talking about “de novo sequencing”, but that is only needed for new organisms types.
E.g. On the image below you can see the reference genome used as a pattern to match against. All the tiny sequenced chunks are attempted to be mapped to the reference, at least partially, to find their best possible position. Repeat it enough times, and there will be a strong guarantee that both all positions are covered, and also that some error rate is tolerated.
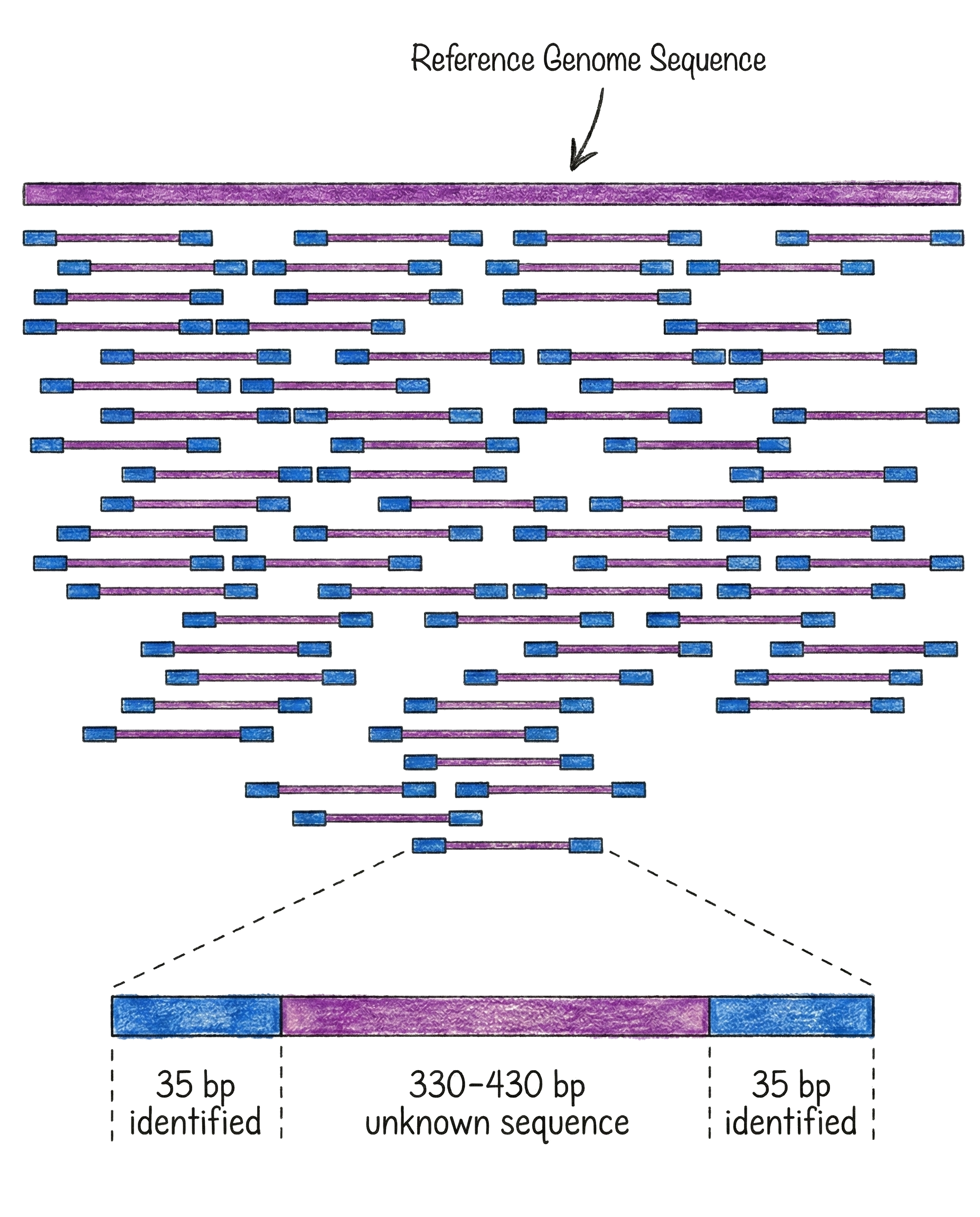
Does this mean that the machine, the methodology and even the lab that does the whole genome sequencing is not really relevant, as long as the coverage is high enough? Unfortunately not.
- Whilst most of the human genome is fairly unique, about 8% of it has a very high repetition count, making the tiny chunks too ambiguous. Long reads are required to fill those gaps4.
- Preparing the DNA for sequencing (library preparation) is usually done via “Polymerase Chain Reaction” for fast and cheap amplification. That process also prefers some regions vs. others and introduces errors. To minimize errors, a more expensive PCR-free preparation is needed.
- Most labs are providing “research” grade whole genome sequencing. That is a fancy way of saying they use PCR protocols and do not have CLIA5 certifications (i.e. lab sterility guarantees) and CAP6 accreditation (i.e. regular inspections), forbidding using their results in clinical settings due to high risk of errors.
Sequencing provider landscape
Whilst there are many sequencing laboratories, they have not been created equal. And even with their significant count, it’s harder than expected to get the procedure done.
To highlight the elephant in the room, offering whole genome sequencing as a service is - simply put - a bad entrepreneurial move.
- The necessary biochemical and clinical knowledge is very high.
- The equipment up-front and maintenance costs are extremely high.
- The reagents for the protocol and human labor costs are very high.
- Customer retention is almost zero, as a genome is sequenced for life.
Because of the above, laboratories employ various tactics that make it very hard for average users to have access to these procedures.
High profile labs which have the capability to do accurate sequencing and also have the capacity to do it in a timely manner will both charge a lot, and will only work with clinics or medical universities as clients. That is their guarantee for recurring charges.
A tier lower on the provider spectrum are labs that work with clinics that allow public sequencing requests. These usually have waiting times ranging from many weeks to many months. These labs try to make their money off of medical grade DNA analytics services, and they will attempt to retain the sequenced genome instead of giving it out to the client. This ensures their subsequent charges when new requests are made against the dataset. To add insult to injury, you will also find gems like below in their terms:

At the bottom of the food chain are direct-to-consumer genomic providers. These are all your usual suspects that are a Google search away. They are as shady as they get.
- They employ extremely aggressive online marketing campaigns while also actively monitoring and censoring dissatisfied customers.7
- They charge subscription fees to have access to your already sequenced data and to provide “ongoing insights” as new “discoveries are made”.
- They emphasize that their products are not clinical grade, and cannot be used for diagnostic or medical purposes.
- Most of them require you to sign away the rights to your genetic data for “research” purposes with “partnering institutes”.
- Some of them lost their genetic data to hackers8, others sold all their user data to venture capital9.
The picture is quite bleak. As a plain person, your only direct choice is a set of exploitative companies that promise everything, deliver “something” and guarantee nothing.
So, what now?
That is a very good question! Our genome has the potential to unlock a lot of interesting secrets, but will probably lead to even more questions than we have now!
Unfortunately, there’s no clear-cut solution on what the next steps are. Of course, before you can do anything with your genetic data, you first have to obtain it, one way or another. To avoid playing king-maker, we will not suggest any specific provider, rather it’s time to do your own homework.
What we would suggest, is:
- Only contract labs offering clinical grade whole genome sequencing.
- Only contract labs having CLIA certification and CAP accreditation.
- Ideally contract local certified medical providers (i.e. hospital).
- Ideally be in touch with professionals for clinical interpretation.
- Optionally collaborate with research institutes (i.e. universities).
- Optionally find, join and help open source bioinformatics projects.
Most importantly, always read the terms-of-service for any provider before sharing one of your most precious personal data (when in doubt, avoid). Do spare the time to investigate each provider in detail and do not FOMO10 in due to limited-time offers. You didn’t know your genetic code until now, a few more days, or weeks, or even months doesn’t matter.
Lastly, have fun, otherwise there’s no point :)
Disclaimer
As a mandatory note to our readers, please be aware, that none of the past, present and future content posted here constitutes medical advice.
Exploring your genetic information, biometric markers or any other health data can be a fulfilling journey, but it will also require a lot of time, effort and money to pursue. As it might not get you the insights you are looking for, only invest as much as you can afford.
Understanding your body and health is an important first step, but please seek out medical advice before acting on your learnt information.
Memo
This post was originally written by Peter on 18th April 2024 for another audience and was lifted into the Dark Bio blog.
Until next time…
-
In certain cases it can happen that an organism has multiple sets of genes, a phenomenon called chimerism, or human chimerism for people. ↩︎
-
You can learn more via a talk by Jonathan Pettitt, titled What Darwin won’t tell you about evolution. ↩︎
-
The different methods are based on different biochemical phenomena. If interested, Sagar Aryal has a nice summary written up. ↩︎
-
For exact details and how the gaps were overcome “de novo” (in 2022!), please see Science: The complete sequence of a human genome. ↩︎
-
CAP: College of American Pathologists laboratory accreditation. ↩︎
-
Dante Labs, for example, was referencing all 5-star reviews, while the true customer feedback on the site they actually quote is 2.2 stars. ↩︎
-
The Guardian: 23andMe admits hackers accessed DNA data of 7m users. ↩︎
-
TechCrunch: Francisco Partners is acquiring MyHeritage. ↩︎
-
FOMO: Fear of missing out. ↩︎
